Software
( GitHub site )
Bayesian Estimation of Non-linear Data (BEND)
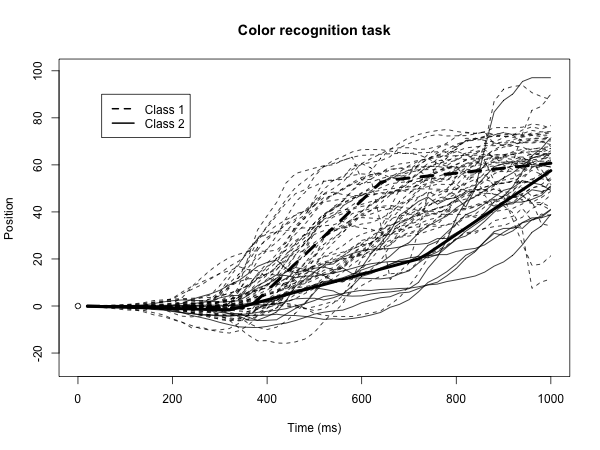
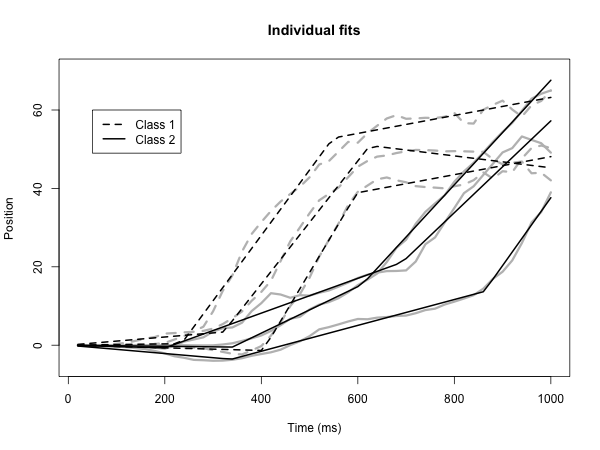
BEND is an R package available on CRAN and GitHub, with a suite of methods to infer Bayesian piecewise growth mixture models for longitudinal data, developed in collaboration with Corissa Rohloff and Nidhi Kohli. Methodology incorporated into the package was published in several articles from 2018-2025, as detailed in the package description. Previously I had developed the Bayesian PGMM package, with more limited functionality, to perform the methodology in this article (Psychometrika, PMID: 29150814, 2018).
Multiway Regression
MultiwayRegression is an R package to predict one multi-way dataset (i.e., tensor) from another multi-way dataset, as described in this article (Journal of Compuational and Graphical Statistics, 2018). The package is available via CRAN at this link and via GitHub at this link.
Multiway Classification
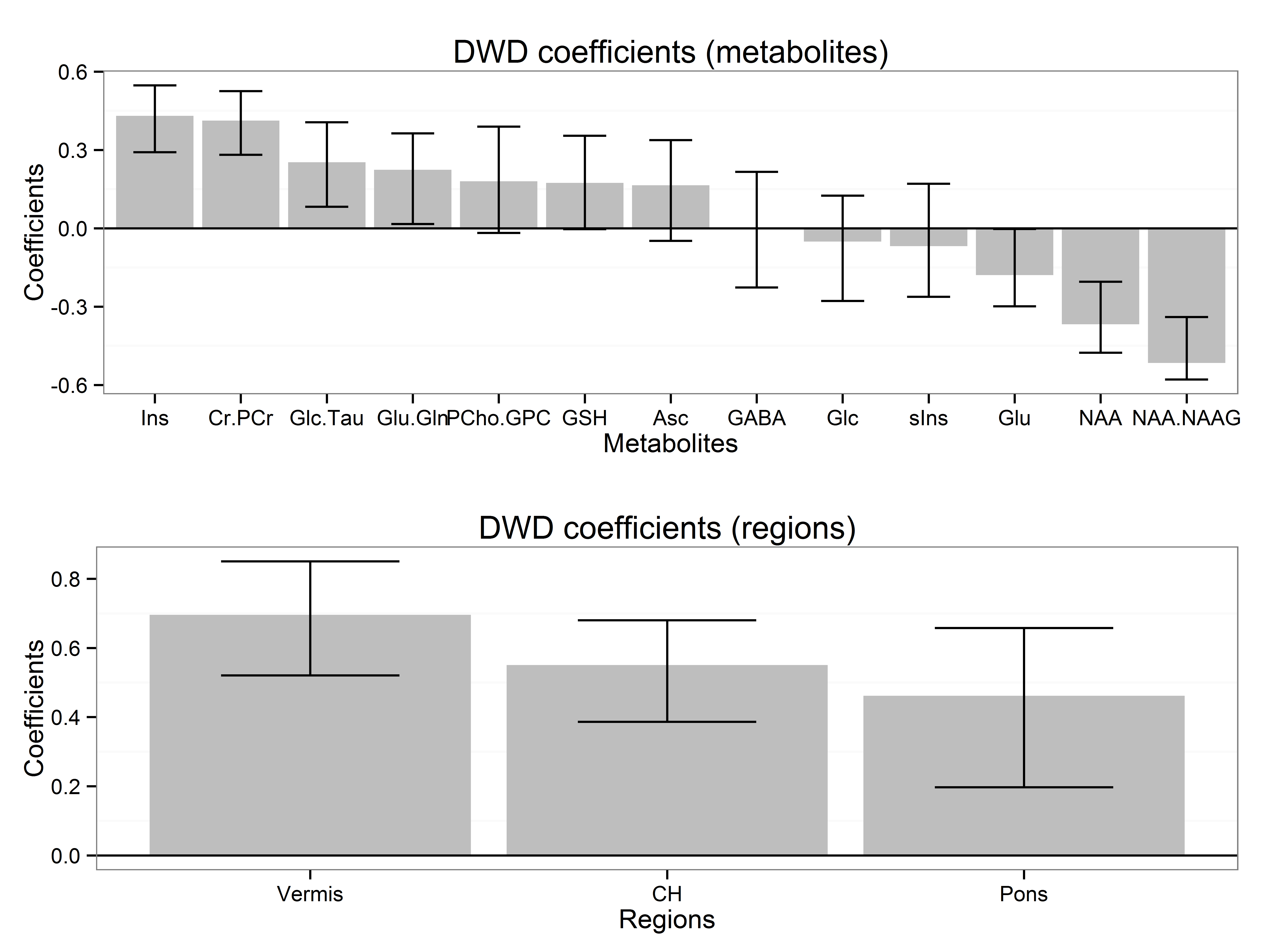
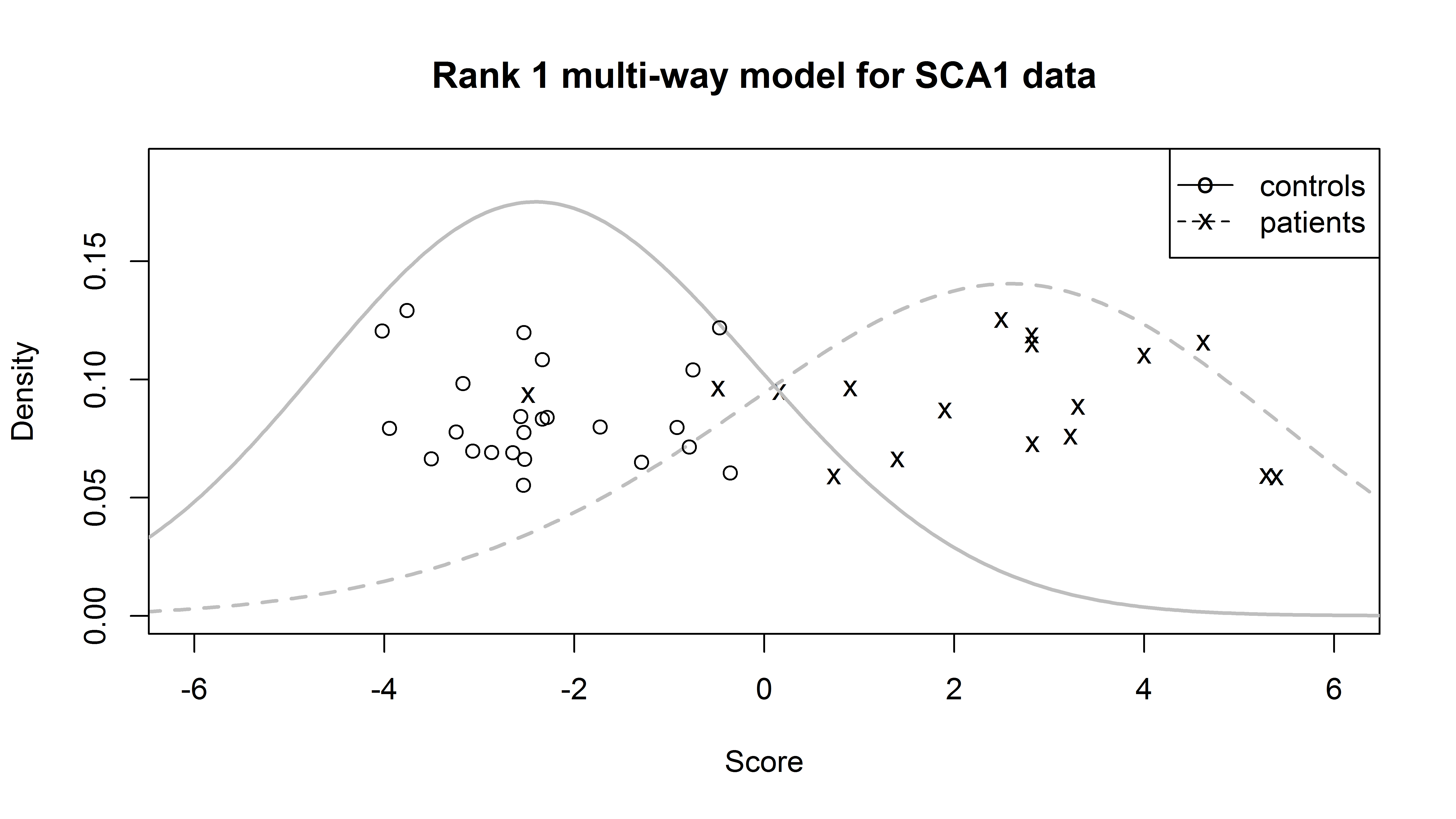
MultiwayClassification is an R package to perform linear classification for data with multi-way structure, as described in this article (Biostatistics, PMID:28115314, 2017). This is joint work with Tianmeng Lyu and Lynn Eberly. The package is available via GitHub at this link.
Bayesian Screening
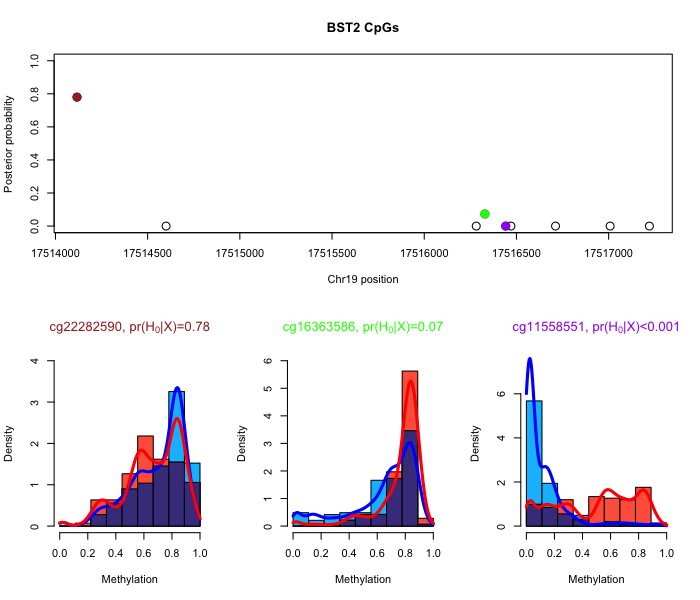
BayesianScreening is an R package to compute posteriors from a gene-level hierarchical prior for genome-wide and epigenome-wide association studies, as described inthis article (Bioinformatics, PMID:31651034, 2020) and this article (Biometrics, PMID:28083869, 2017). Also includes functions to perform two-class testing with shared kernels for methylation array data (see this article in Biometrika). This is joint work with David Dunson. The package is available via GitHub at this link.
Bayesian Consensus Clustering
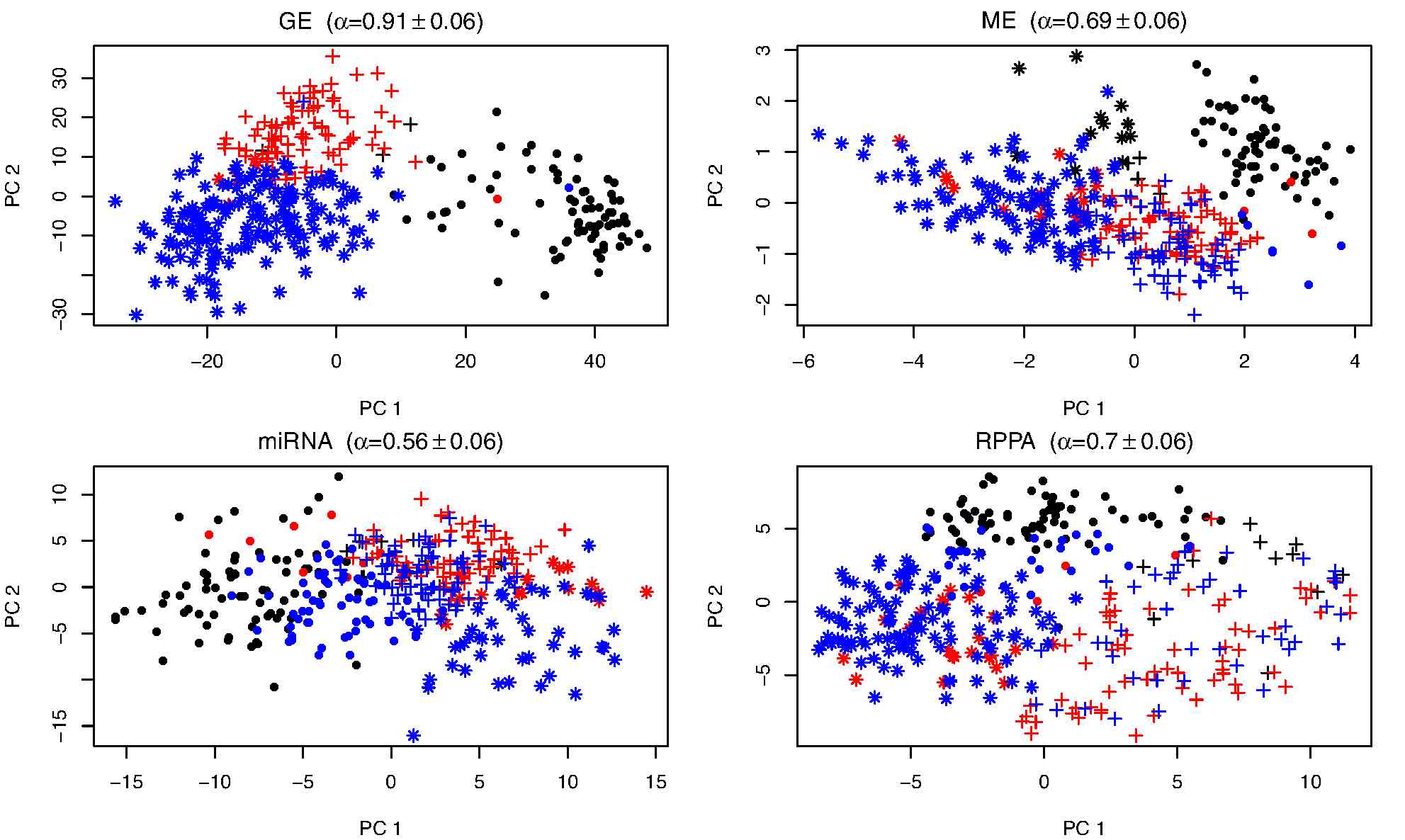
Bayesian consensus clustering is a tool to cluster a set of objects based on multiple sources of data. The model permits a separate clustering of the objects for each data source that adhere loosely to an overall clustering. The method is described in this Bioinformatics article, and this zip folder contains R code with instructions and examples. This is joint work with David Dunson. A user friendly R package to perform BCC, developed and maintained by Tim Triche, is available at this link.
Joint and Individual Variation Explained (JIVE)
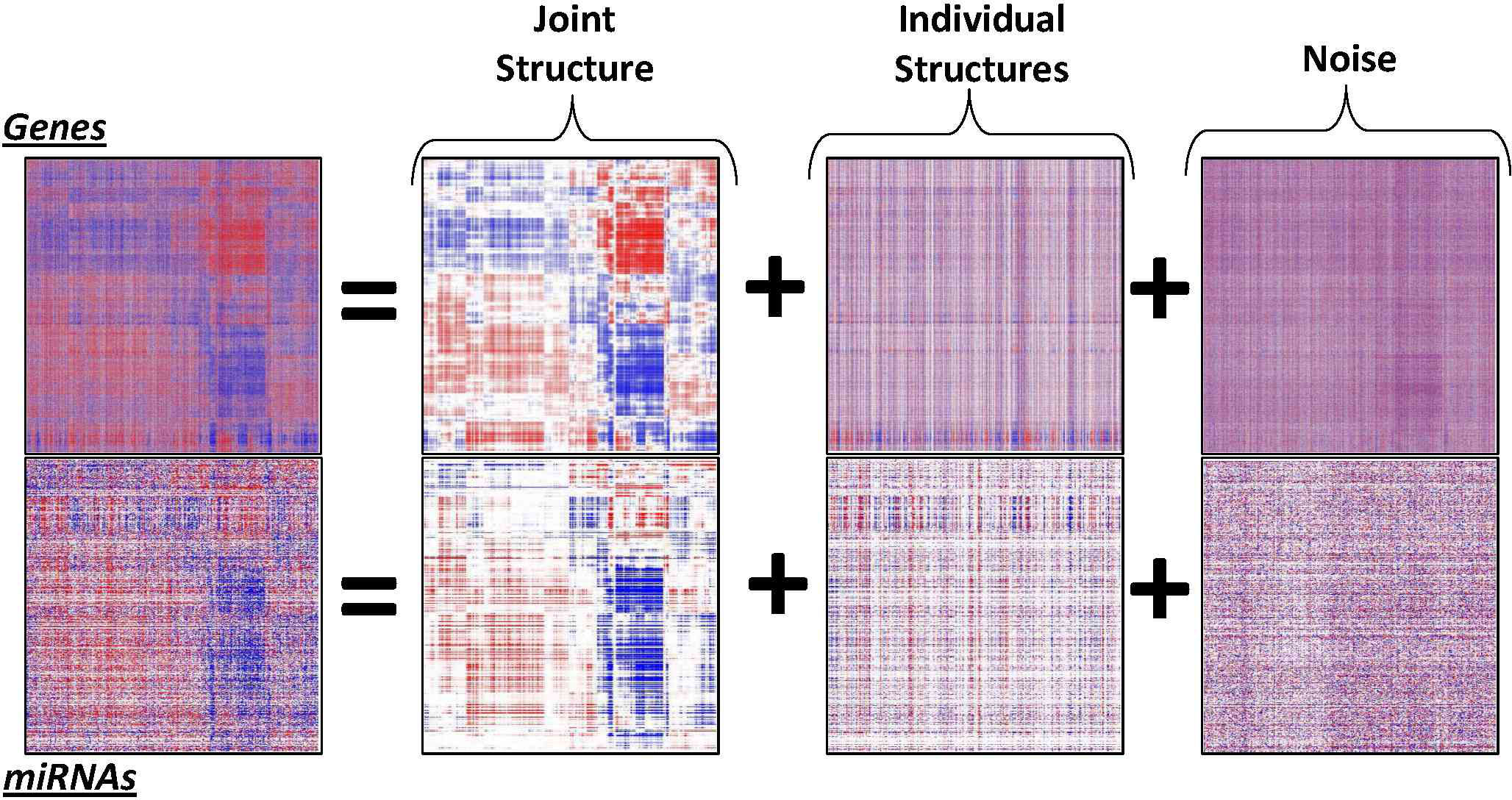
JIVE is a flexible exploratory method for the integrated dimension reduction and visualization of multiple datatypes on the same set of samples. Original Matlab scripts and sample data are available here, and a user-friendly R package with some enhancements is available here. The original manuscript is published in the Annals of Applied Statistics (link), and a the R package is described in Bioinformatics (link) with enhancements to predict an outcome described here (2017) and here (2022). The original development was joint work with Katherine Hoadley, Steve Marron, and Andrew Nobel; the R package was developed with Michael O'Connell, Adam Kaplan, and Elise Palzer.
Primer Efficiency Analysis
A collection of statistical methods to analyze the efficiency of a set of primer-pairs for quantitative real-time PCR. The R script PEA.r contains code to provide individual efficiency estimates with confidence, identify and remove unreliable primers, cluster amplification efficiencies, and adjust CT values. See the PEA user's guide for setup instructions, function descriptions and illustrative examples. The related manuscript is published in BMC Bioinformatics (link). This is joint work with Dirk Dittmer.
Binary Biclustering
Matlab code to bicluster a binary data matrix can be found here. This code is adapted from the LAS method, and uses a binomial score function. For more details, see this short report.
Classification Based Biclustering
Matlab code to search for biclusters that are distinguishing between two sample classes can be found here.
For more details, see this short report. (This code gives a preliminary approach and has not been well tested - use at your own risk.) .
ToxPi GUI
ToxPi is a flexible tool to prioritize environmental chemicals based on diverse toxicity data. Developed in collaboration with David Reif, Myroslav Sypa, Ivan Rusyn, Fred Wright and others.
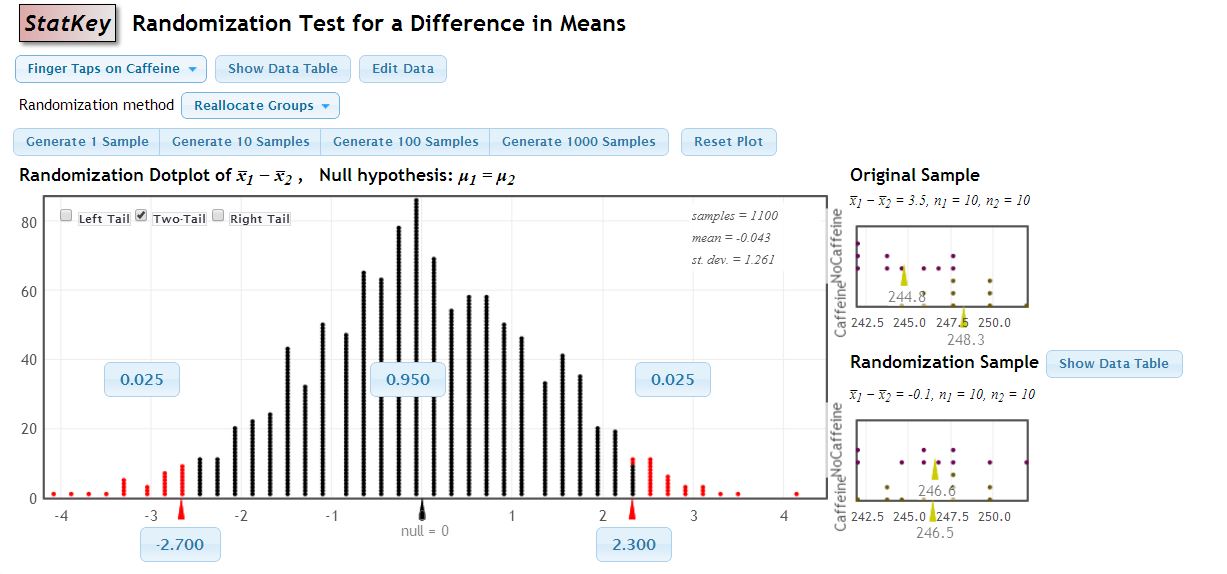
StatKey
StatKey is a collection of easy-to-use online applets to visualize bootstrapping and randomization tests. Also includes online applets for descriptive statistics and theoretical probability distributions. Developed in collaboration with Rich Sharp, Ed Harcourt, Kevin Angstadt, Patti Frazer Lock, Robin Lock, Kari Lock Morgan and Dennis Lock.